All talk and poster papers are available for download at http://purl.org/ronco/swat The organizers would like to have everyone’s suggestions for future conferences, and also for topics and tasks for the hackathon tomorrow.
Keynote by Dov Greenbaum
This is a talk about the legal issues that come with FAIR. The Zvi Meitar Institute focuses on examining the ethical, legal and social implications of new and emerging technologies. Examples include fake news and space law.
Big data: velocity, veracity, variety, volume. And a reminder that if something is free, you’re not the customer, you’re the product. The Hague declaration provides some nice numbers about academic data. By 2020, the expectation is that there will be 44 zettabytes, with 90% being generated in the last 2 years. 2.4 million scholarly articles publised in 2013, or one every 13 seconds. It’s estimated that big data is worth more than $300 billion per year to companies. UNESCO (specifically, the International Bioethics Commitee or IBC) is worried about how data is used. Health data comes from many sources: home monitoring, EHR, lab records, etc.
The IBC has a report from 2017 that asked: Do you own your own data? Is it yours to give? What about ownership and possession?
Now, thinking about this in the context of FAIR, particularly the accessible and reusable parts, which leads to a discussion of licensing. Open source licensing has one problem: in order to license data, you must own it legally. The way we own non-tangible items is mostly through intellectual property law. IP includes patents, copyright, trademark, trade secret, and sui generis (a new form of IP law). In 1916, a uk ruling stated that “the rough practical test that what is worth copying is prima facie worth protecting”. IP rights do not grant you the right to do anything, they just allow you to grant permission – a negative right rather than a positive right.
Is the big data patentable? Then you can license it… Otherwise we need to find another way (see IP methods above). To be patentable it must be patentable subject matter, utility (some kind of use, and can’t be “evil”, or a tax cheat – utility in the US is a very weak definition – allows virtually anything to have utility), novel, non-obvious. But Big Data doesn’t fit into the patent definition of “a useful process, machine, manufacture, or composition of matter”. This means also that wild plants, DNA, algorithm, laws of nature are unpatentable. In short, Big Data is not patentable subject matter.
Next form of IP law is copyright. This covers original works of authorship fixed in any tangible medium of expression – this includes things like databases. Copyright is automatic and free (unless you want to register it), unlike patents. If you are using AI to create some of your data, you probably don’t have authorship. What is copyrightable? literary works, musical works, dramatic works, choreographed works, pictorial graphic sculptural sound works, architectural works. Ideas, laws, facts, discoveries are all not copyrightable. Data is facts, and therefore not copyrightable. Copyright protects your expression of an idea, not the idea itself. You cannot copyright the building blocks of knowledge – to promote innovation. There was a supreme court case in 1991 that centered around the yellow pages – supreme court decided that there would be no more protection of databases of facts. However, they said you could protect databases if you had a contract and the other side agreed to the contract, you could copyright your data. The contract for some software would be as soon as you open a CD of software, or a EULA online, the contract is implicit. The AWS contract mentions a zombie apocalypse 🙂 (clause 47 I think I heard?!) If you have a factual database, how can you prove that someone has copied your data? The easiest way is to salt – dictionaries have fake words, google maps has fake cities, phone books have fake people etc. In short copyright is no good.
Next form of IP law is trade secrets. This is fine, but you want to be FAIR and trade secrets are the opposite of FAIR. So, if you don’t own big data under IP, how do you license it?
WTO says that you should be able to protect computer programs and compilations of code, e.g. the database itself if not the data. Most countries have not incorporated this yet. US has the digital millennium copyright act, but it’s a weird kind of IP right – you can claim ownership of your databases only if you protect it with a digital rights management tool. Then when someone has hacked into you, they have infringed on your rights (kind of backwards really). This is most prominent in DVDs and violating the region protection. The EU database directive contains database protection (1996). They were thinking mostly of betting databases! This is sui generis – a new type of IP protection for databases. However, it really only protects the structure of the database and not the content, and you have to have made a substantial investment and effort. In 2018 an analysis was performed, and academics were very unhappy with the directive as they weren’t sure what classified as violations of the law, and 2/3 thought it had a negative effect – the sui generis right was considered to have a negative effect on legal certainty.
How do you protect data right now? You contract it or you use antitrust law. Ownership grants you benefits and control over that information, but possession does not mean ownership. Possessing data doesn’t mean you own the data. The Hague Declaration is something you can “sign” if you agree, and tries to find solutions to this.
So, what we need is a complete change. We need to think about multiple owners: the patient, the doctor, the informatician. Perhaps we can’t find an ownership right in data – so perhaps we should look at it as having custody rather than ownership of the data. Custody brings liabilities (e.g. GDPR) and responsibilities. Custody also means good stewardship, privacy, security, attribution, FAIR principles. Individuals could transfer some of their rights via informed consent. But this isn’t a full solution, and this is still an ongoing problem.
Q: A lot of life science databases use CC BY, CC BY-SA – is this correct?
A: This is not legal advice. In Europe, via the database directive, databases are protected so if you take the entire databases you are infringing. If you extract data without maintaining the structure of the database, you are not infringing. This is one reason why academics are unsatisfied with the EU database directive.
Q: It’s a creative act to map data to e.g. ontologies. Is this enough to be able to have it be an act of creation and makes it protectable?
A: Modified data that is still factual (e.g. cleaning data, adding ontology terms) does not change the fact that it’s factual – and therefore cannot be protected. Might still fall under the database directive, as described above.
Q: Patients possess (not own) their data. In MS research, patient groups take custody of that data. Is this style the future direction?
A: It makes sense to get informed consent from a group – the group could use a trade secret type that requires an NDA to use, for example.
Documentation Gap in Ontology Creation: Insights into the Reality of Knowledge Formalization in a Life Science Company
Marius Michaelis (speaker) and Olga Streibel, Bayer
For them, ontology creation begins with specification / competency questions, then conceptualization of the ontology via use cases, implementation into a formal knowledge mode (ontology), then ontology testing. He will concentrate on the conceptualization phase.
Within the conceptualization phase, you have knowledge acquisition by knowledge engineers who, on the one hand, research explicit knowledge, and on the other hand also elicit tacit knowledge. This process of eliciting knowledge takes a lot of effort and is time intensive. How can you document this stage? Such documentation would encourage collaboration, prevent knowledge loss, and encourage clarity.
Within Bayer, they ran a survey of 13 knowledge engineers and 16 domain experts. He will discuss both the timepoint and nature of how knowledge engineers document. Most start documenting while or after the model is being created, which means during implementation rather than during conceptualization. The respondents also had a roughly equal mixture of structured and unstructured methods (don’t necessarily follow guidelines). But what we want is joint, timely, structured documentation.
His bachelor’s thesis (University of applied sciences Potsdam), “Documentation concept for the exchange of knowledge in the process of creating ontological knowledge models.”
Partitioning of BioPortal Ontologies: An Empirical Study
Alsayed Algergawy and Birgitta König-Ries
BioPortal, AgroPorta, OntoBee, and EcoPortal (Allyson: EcoPortal apparently last updated 2 years ago?) all exist to store publicly-available ontologies.
Most existing studies of bioportal ontologies focus on ontology reuse and ontology evaluation (the quality of the ontology). A few also consider the partitionability/modularization of ontologies (e.g. ISWC 2011). They also looked at the partitioning of BioPortal records.
Overall workflow: 1. get ontologies 2. transform into owl/obo 3. partition 4. analyse. Some existing partitioning tools include OAPT and PATO(?, not *that* PATO I suppose). They developed OAPT, which has these steps for partitioning: 1. ranking ontology concepts 2. determine cluster heads 3. partition 4 generate modules. In addition, in Manchester they developed a tool called AD (Atomic decomposition). So, they applied AD, OAPT and PATO to BioPortal’s 792 ontologies. For details, see https://github.com/fusion-jena/BioPortal-Ontology-Partitioning.
They discussed how many modules were created within each ontologies – you can specify in OAPT the optimal number of partitions. There were three 0-module ontologies (SEQ, SMASH, ENM). Both 0-module and 1-module ontologies don’t seem to be fully developed. 100 ontologies generated 2 modules. Over half of the accessible ontologies (347) can be partitioned into only 5 modules. Most of the ontologies which failed to partition seemed to be because of some particular characteristics of the ontologies rather than the tools themselves.
Annotation of existing databases using Semantic Web technologies: making data more FAIR
Johan van Soest (speaker), Ananya Choudhury, Nikhil Gaikwad, Matthijs Sloep, Michel Dumontier and Andre Dekker
They have hospital data that remains in the hospital together with the analysis, and then they submit results to a central location. This is good for patient privacy but not for researchers wishing to access the data. Therefore it relies on good semantic tagging.
Not all hospitals are academic hospitals, and therefore might not have the resources to add the data system to it. So they provided the hospitals with a tool that separates structure from the terminology conversion – this allows the IT person to do the conversion and the domain expert to do the terminology mapping (R2RML). This works but is high maintenance, so they’ve changed the approach. Instead, keep the original data structure and annotate the existing schema.
Their use case was a set of 4000 rectal cancer patient data and used Ontotext GraphDB 8.4.1. They had an ontology with 9 equivalent classes and 13 value mappings (e.g. “m” means male”). Two parent classes for each – the ontology class and the SQL table column.
They are only annotating existing data sources – although they are using non-standard (local) schemas, it does mean there would be no data loss upon any conversion and also they don’t have to make big changes to local systems. Keeping local systems also means that there is a smaller learning curve for the local IT team. They would like to develop a UI that would hide the “ontology-ness” of their work from their users.
FAIRness of openEHR Archetypes and Templates
Caroline Bönisch, Anneka Sargeant, Antje Wulff, Marcel Parciak, Christian R Bauer and Ulrich Sax
HiGHmed was begun with the aim to improve medical research and patient care, and to make data from research and patient care accessible and exchangeable. The project has a number of locations across Germany. Part of this involves the development of an open interoperable and research-compatible eHealth platform to support local and cross-institution patient care and research.
openEHR is a virtual community for transmitting physical health data in electronic form. Archetypes are aggregated into Templates, which then are published and versioned via the CKM (Clinical Knowledge Manager). They have assessed their archetypes and principles in the context of the FAIR principles, and found that they were compliant with 13/15 of the principles.
A Working Model for Data Integration of Occupation, Function and Health
Anil Adisesh (speaker), Mohammad Sadnan Al Manir, Hongchang Bao and Christopher Baker
The Occupational Heath Paradigm is a triangle of work-illness-wellness. In Canada they have a NOC Career Handbook with 16 attributes that help define the requirements of various careers. This can be helpful when someone with an injury wishes to change to a different job, but one that is similar enough to their previous job that there isn’t a lot of retraining. A semantic model is populated with coded patient data representing disease (ICD-11), functional impairment (ICF), occupation (NOC), and job attributes (NOC Career Handbook). The NOC coding for the data was done manually initially, and then they developed a coding algorithm to assign the occupations automatically. The algorithm starts with a textual job title and then progresses through a number of steps to get a NOC. They use sparql queries and semantic mapping to suggest job transitions to accommodate a functional impairment.
They did some test queries with their model to see if they could classify people in jobs according to their attributes, e.g. if a person has disease J62_8 them what jobs could they do? What job with a patient with visual impairment likely return to (previous job + impairment = new job options)?
This work could be applicable in finding work for immigrants and newcomers, finding comparable work for people with an acquired disability, or people with accidental injuries that could otherwise end up on long-term disability. The model seems fit for purpose to integrate info about occupational functioning and health.
FAIR quantitative imaging in oncology: how Semantic Web and Ontologies will support reproducible science
Alberto Traverso (speaker), Zhenwei Shi, Leonard Wee and Andre Dekker
Medical images are more than pictures, they are big data. Indeed they are the unexplored big data as many images are stored and not re-used, as well as having more information than is used in the first place. There were 153 exabytes in 2013, and 2,314 exabyte expected in 2020. Within medical imaging, the combination of the image and “AI” results in what is called quantitative imaging. Machine learning is used to create prediction models (e.g. the probability of developing a tumor, or a second tumor).
There is currently no widespread application of “radiomics” technology in the clinic. The data we produce grows much faster than what they can currently do with the models. Radiomics studies lack reproducibility of results. The challenges and some solutions are: models work but only on their own data (fix with multi-centre validation); a lack of data-driven evidence and external validation (fix with TRIPOD-IV models); lack of reproducibility (fix with sharing of metadata in a privacy-preserving way); explosion of computational packages; how can models be shared when data isn’t; poor quality of reporting; lack of standards; a need for domain knowledge (these last three can be fixed by standardized reporting guidelines, increased agreement, and data-driven meta analyses). FAIR quantitative imaging = AI + medical imaging + open science.
“Ontologies function like a brain: the work and reason with concepts and relationships in ways that are close to the way humans perceive interlinked concepts.”
Radiomics Ontology (RO) – https://github.com/albytrav/RadiomicsOntologyIBSI/wiki/1.-Home
Appropriate questions for this system: What is the probability of having esophagitis after irradiation in certain patients that received a dose of…? What is the value of Feature X for rectal cancer patients with a poor prognosis computed on an ADC scan?
Keynote by Helen Parkinson
The Odyssey has a non-linear plot, and Helen is using the monsters and challenges to hang her topics off of.
Helen asked the question in August: if there are no FAIR-compliant vocabularies, how can you be FAIR? If there aren’t any, then the FAIR indicator cannot be satisfied. Therefore you have a recursive, circular issue 🙂
Why do we need to be FAIR? What tools do we need to be FAIR? How do we know we are FAIR? I2 of FAIR is to use vocabularies that adhere to FAIR principles. EBI is at 273 PB of storage, with 64 million daily web requests in 2018. As with metadata in many projects, the metadata is often troublesome. They would like to build FAIR capable resources (but we’re not quite sure what FAIR capable is yet); acquire, integrate, analyse and present FAIR archival data and knowledgebases (and all the resources are very different – context is important); determine the FAIR context for our resources and data; define what it means to be a FAIR vocabulary.
Within the SPOT group, they develop a number of ontology applications, e.g. that make use of the data use ontology. For Helen, there are temptations associated with formalisms – the more complex the ontology, the more time/money it will cost but you will get a strong semantics. http://FAIRassist.org provides a list of current FAIR tools.
Which features of a FAIR vocabulary are already defined in the OBO Foundry? Many are already aligned, but some parts of the Foundry are deliberately not aligned, including: open, defined scope, commitment to collaboration, reused relations from RO, and naming conventions. These are parts of the Foundry that are not, and probably should not, be a required feature of a FAIR vocabulary. So then they mapped the aligned part of the foundry to the FAIR principles.
When talking about “deafness”, we need to consider the assay, the disease and the phenotype – and they all need to be connected – making all this interoperate is important. To help, they developed OXO which provides cross-references between ontologies, but it doesn’t tell you anything about the semantics.
FAIR Vocabulary Features- required (from Helen’s POV)
- Vocabulary terms are assigned globally unique and persistent identifiers plus provenance and versioning information
- Vocabularies and their terms are registered or indexed in a searchable resource
- Vocabularies and their terms are retrievable using a standardised communications protocol
- the protocol is open, free, and universally implementable
- Vocabularies and their terms persistent over time and appropriately versioned
- Vocabularies and their terms use a formal accessible and broadly applicable language for knowledge representation
- Vocabularies and their terms use qualified references to other vocabs
- released with a clear and accessible data usage licence
- include terms from other vocabs – when this happens, import standards should be used.
- meet domain relevant community standards.
Why should vocabs be FAIR? Ontologies are data too, and data should be FAIR. How do we know we are FAIR? When the vocab has use, reuse, implementation, and convergence.
Where is FAIR in the Gartner Research’s Hype Cycle? 🙂
Q: should FAIRness be transitive? Should FAIR vocabs only import FAIR vocabs?
A: I would like it to be, but it probably can’t always be.
The OHDSI community has to deal with this issue of transitive FAIRness already. Sometimes they import only certain versions of the ontology. They don’t think it’s possible/practical to move the entire healthcare industry to FAIR standards.
Q: What would be a killer application for speeding up and making data FAIR?
A: Proper public/private collaborative partnerships. Getting large organizations on board for, at a minimum, a few exemplar organizations. One field might be antimicrobials and another would be neurodegenerative diseases as they are biological difficult and traditional routes of pharma haven’t worked as well as hoped.
Afternoon Panel
This panel is about the conference itself and our thoughts on it. During the panel session, topics included:
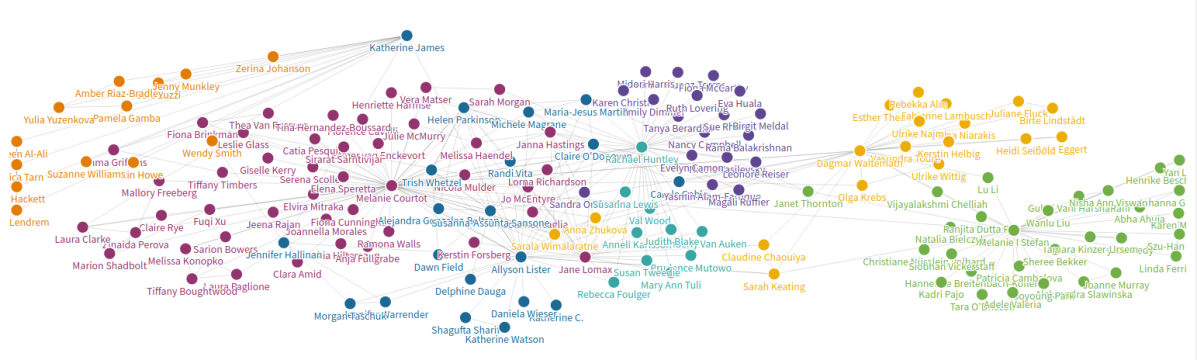
- How can we improve gender diversity in the organizing group? From this, corollaries came up such as making the conference more relevant to other parts of the world, e.g Asia. Equally, this workshop has been organized on a shoestring budget and via people volunteering their time. The question is – what do we want from the conference in the future?
- Do you see food safety as a potential topic for SWAT4HCLS? Yes, but we need to consider what our scope is, and how adding something like “food safety” would impact upon the number of papers submitted. e.g. there has been a big agrisemantics group last year and this, yet the conference name wasn’t changed.
- Tutorial logistics should be improved for next year. The community haven’t submitted many tutorials this year. Should we keep them?
- How do we leverage our knowledge to help the community at large? Should we reach out and bring people in via training?
- The conference has been going on for over a decade might some kind of retrospective be appropriate? Perhaps combine it with a change in direction, said one panel member. Perhaps someone present something lighthearted for next year and present the changes?
- Should we have a student session? Well, we already have some of them, even presenting already at the conference. We should work more to get students from the local university to participate, as they haven’t really taken up the offer in the past.
- Should we remove the HC, because we continue to expand into other areas and can’t keep adding letters to the name! If so, what should the name be? Probably shouldn’t change it.
- Where is it going to be next year? They can’t tell you yet.
- Should we invite a big hitter from a neighbouring area to pull in more communities? Should we include expand to end users?
Please also see the full programme. Please note that this post is merely my notes on the presentations. I may have made mistakes: these notes are not guaranteed to be correct. Unless explicitly stated, they represent neither my opinions nor the opinions of my employers. Any errors you can assume to be mine and not the speaker’s. I’m happy to correct any issues you may spot – just let me know!







![Because tardigrades are lovely animals, taxonomically speaking of course. Aditya Sainiarya [CC BY-SA 3.0 (https://creativecommons.org/licenses/by-sa/3.0)]](https://themindwobbles.files.wordpress.com/2019/12/water_bear.jpg?w=1061)
You must be logged in to post a comment.